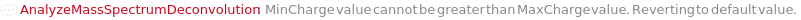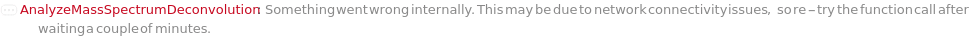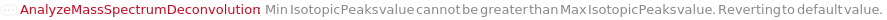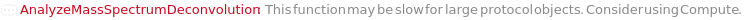AnalyzeMassSpectrumDeconvolution
AnalyzeMassSpectrumDeconvolution[Data]⟹Object
denoises and centroids spectral data, and then combines peaks corresponding to analytes of the same elemental composition, but different isotopic compositions, into monoisotopic peaks. Optionally, peaks with known charge states can be shifted to the corresponding single-charged position.
AnalyzeMassSpectrumDeconvolution[Protocol]⟹Object
denoises and centroids spectral data in the linked objects of the Data field of the Protocol, and then combines peaks corresponding to analytes of the same elemental composition, but different isotopic compositions, into monoisotopic peaks. Optionally, peaks with known charge states can be shifted to the corresponding single-charged position.
Details
- The purpose of this function is to detect and combine the peaks of isotopic clusters in mass spectrum data, and generate a spectrum with reduced complexity for use in additional analyses, such as spectral annotation, with increased effectiveness. Isotopic clusters, which are set of peaks corresponding to analytes of the same elemental composition, but different isotopic compositions, are identified via an algorithm with requires no prior knowledge of the analytes. First, the algorithm removes noise and creates a list of centroided peaks. Then, starting from the lowest m/z value, the algorithm scans each peak in the spectrum, and for each peak, checks if the position is within some distance of the previous peak. If the option UseDecreasingModel is set to True, the algorithm also checks that the intensity of the peak is less than the intensity of the previous peak. If a peak satisfies both the position and intensity criteria relative to the previous peak, the peak is added to the isotopic cluster. This process continues until some peak does not satisfy the criteria, at which point, the algorithm will begin assigning peaks to a new isotopic cluster. Once the algorithm has successfully identified all the peaks of an isotopic cluster, it creates a monoisotopic peak in the deconvoluted spectrum corresponding to that cluster. More detail can be found in the following paper: https://pubs.acs.org/doi/full/10.1021/acs.jproteome.0c00544.
-
ChargeRange MinCharge value cannot be greater than MaxCharge value. Reverting to default value. FatalError Something went wrong internally. This may be due to network connectivity issues, so re-try the function call after waiting a couple of minutes. IsotopicPeaksRange Min IsotopicPeaks value cannot be greater than Max IsotopicPeaks value. Reverting to default value. MassSpectrometryObject This function may be slow for MassSpectrometry data objects which do not yet contain an MzMLFile. Consider using Compute. The first time this function is run, an MzMLFile will be added to the data object, allowing for much faster analysis in the future. OutputOption AnalyzeMassSpectrumDeconvolution does not support the Output options Preview and Result simultaneously; Must use only one at a Time. Output options Options and Tests can be used with either. ProtocolObject This function may be slow for large protocol objects. Consider using Compute. StartIntensityCheck The option StartIntensityCheck has no effect on the results if the option IncludeDecreasingCheck is set to False.
Input

Output

Peak Clustering Options

Post-Processing Options

Pre-Processing Options

Method Options
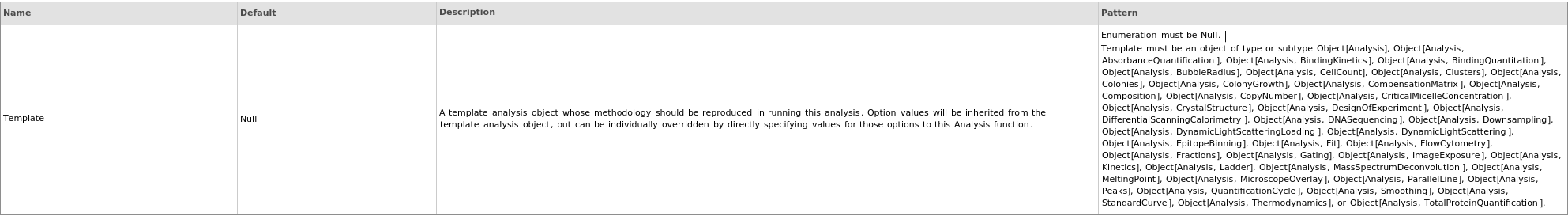
Messages
Examples
open allclose allBasic Examples (4)
Additional Examples (2)
Options (12)
AveragineClustering (1)
ChargeDeconvolution (1)
IntensityThreshold (1)
IsotopicPeakTolerance (1)
KeepOnlyDeisotoped (1)
MaxIsotopicPeaks (1)
MinIsotopicPeaks (1)
SmoothingWidth (1)
StartIntensityCheck (1)
Messages (6)
ProtocolObject (1)
Last modified on Thu 18 Sep 2025 15:08:19