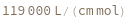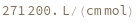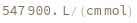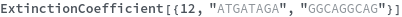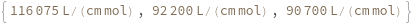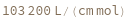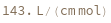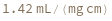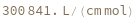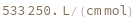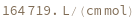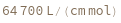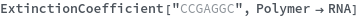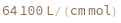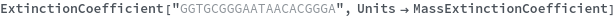ExtinctionCoefficient
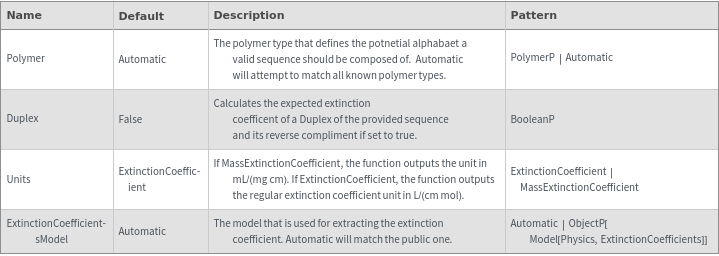
ExtinctionCoefficient[n]⟹extinction
determines the extinction coefficent at 260 (nm) of an average sequence of length n.
ExtinctionCoefficient[sequence]⟹extinction
determines the extinction coefficent at 260 (nm) of the provided sequence.
ExtinctionCoefficient[strand]⟹extinction
determine the extinction coefficent at 260 (nm) of the provided strand.
ExtinctionCoefficient[structure]⟹extinction
determine the extinction coefficent at 260 (nm) of the provided structure.
ExtinctionCoefficient[object]⟹extinction
determine the extinction coefficent at 260 (nm) of the provided object.
Details
- The Extinction Coefficient calculation is based on the formula as described in Object[Report, Literature, "id:pZx9jon4xKW9"]: V. Bloomfield et al, "Nucleic acids: structures, properties and functions." (2000) p174-176.
- For DNA, the extinction coefficient and the hyperchromacity at 260 nm values can be found in Object[Report, Literature, "id:kEJ9mqR5EmJB"]: A. V. Tataurov et al, "Predicting ultraviolet spectrum of single stranded and double stranded deoxyribonucleic acids." (2008) p66-70.
Input
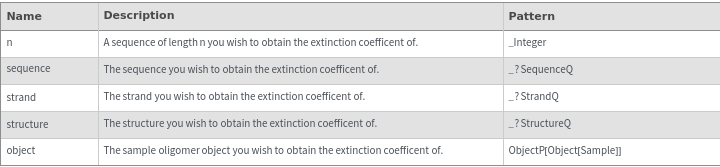
Output

General Options