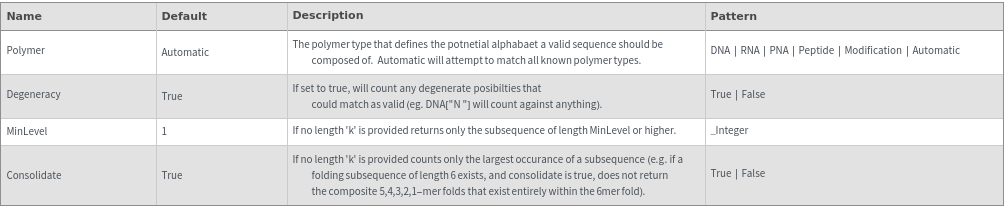
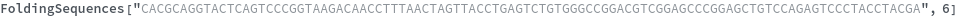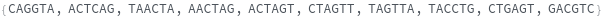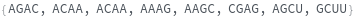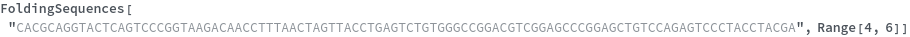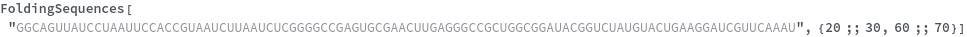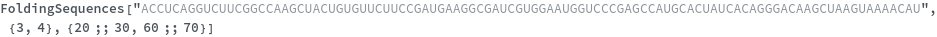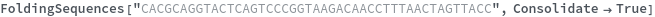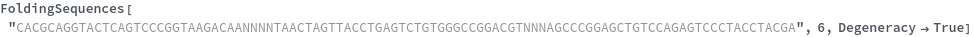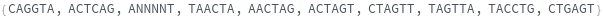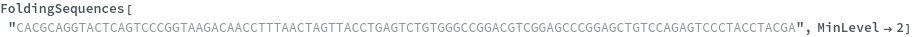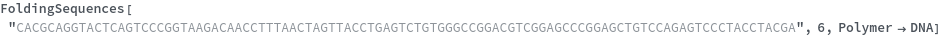FoldingSequences
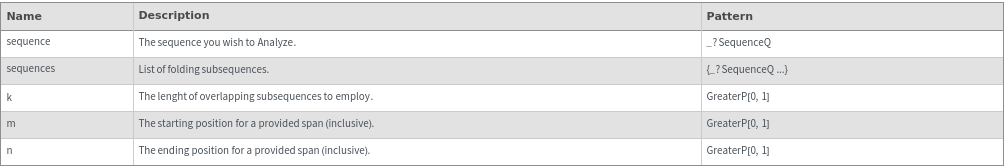
FoldingSequences[sequence,k]⟹sequences
this function takes a sequence and returns a list of subsequences of length k from the entire sequence that could potentially fold.
FoldingSequences[sequence,k,Span[m,n]]⟹sequences
this function takes a sequence and returns a list of subsequences of length k from the position m to positon n in the sequence that could potentially fold.
FoldingSequences[sequence]⟹sequences
this function takes a sequence and returns a list of subsequences from length spesified by the MinLevel option to the largest fold length present from the entire sequence that could potentially fold.
FoldingSequences[sequence,Span[m,n]]⟹sequences
this function takes a sequence and returns a list of subsequences from length spesified by the MinLevel option to the largest fold length present from the position m to positon n in the sequence that could potentially fold.