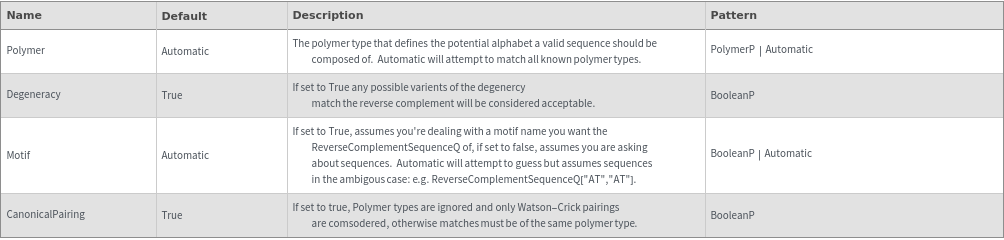
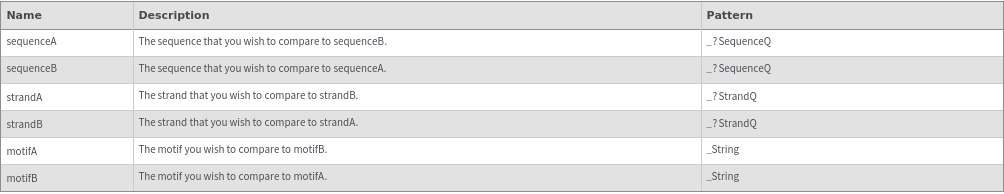
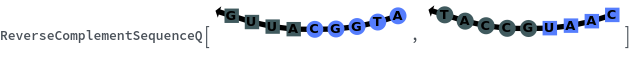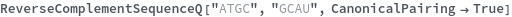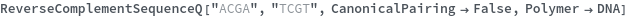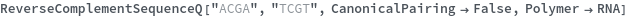ReverseComplementSequenceQ
ReverseComplementSequenceQ[sequenceA,sequenceB]⟹out
returns true of sequenceA is the reverse complement of sequenceB.
ReverseComplementSequenceQ[strandA,strandB]⟹out
returns true of strandA is the reverse complement of strandB.
ReverseComplementSequenceQ[motifA,motifB]⟹out
returns true if motifA is the reverse complement of motifB (name motif with a ' after it).
Examples
Basic Examples (3)
Additional Examples (4)
Options (13)
CanonicalPairing (2)
Degeneracy (4)
The Degeneracy option can be used to allow degenerate sequences to match:
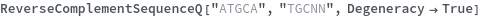

Or if set to False the Degeneracy option can be used to require a strict match:
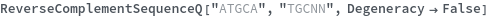

Degenerate numeric sequences are considered matches when Degeneracy is set to True:


Degenerate numeric sequences are not considered matches when Degeneracy is set to False:


Motif (5)
The Motif options is used to disambiguate cases where a the inputs could be sequences or motifs:


If the Motif option is set to False, the function will assume its considering sequences:


If the Motif option is set to True, the function will assume its considering motif names:


If the Motif option is set to Automatic and the case is ambigous, the function assumes its considering sequences:


In the context of a sequence, motif names are ignored by ReverseComplementSequenceQ: