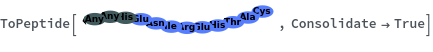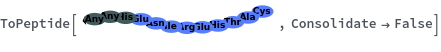ToPeptide
ToPeptide[proteinSeq]⟹peptide
converts the proteinSeq in the single letter amino acid abbreviation format to the peptide the triple letter format.
Examples
Basic Examples (2)
Convert a single implicitly typed protein sequence in the single letter amino acid form into an explicitly typed Peptide in the triple letter abbreviation form:

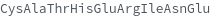
Convert a single explicitly typed protein sequence in the single letter amino acid form into an explicitly typed Peptide in the triple letter abbreviation form: